*by parasites here I am referring to all kinds of infectious disease causing agents including bacteria, viruses, fungi, protozoa, helminths and arthropods.
Why do we care about primate parasites?
Many of the most devastating infectious diseases in humans have origins in wildlife. For example, the global AIDS pandemic originated through human contact with wild African primates and influenza viruses circulate among wild bird populations. These are not only historical occurrences. Recently, for example, rodents were identified as the source of a Hantavirus outbreak in Yosemite National Park, USA . As human populations continue to expand into new areas and global changes in temperature and habitat alter the distributions of wild animals, humans around the world will have greater contact with wildlife. Thus, understanding which infectious agents have the potential to spread from animals to humans is crucial for preventing future human disease outbreaks.
Many efforts are being made to collate information on wildlife and human diseases. Much of my research (which I will blog about when I get chance!) uses an amazing database known as the Global Mammal Parasite Database or GMPD for short. Every time a paper is published which contains details of parasites found in either primates, carnivores or ungulates, the information is added to the database. As much data as possible is recorded, including the species infected, the type of parasite, the prevalence of the parasite, and the geographic location of the study. Prof. Charles Nunn and his colleagues have been collecting data for the GMPD since around 2005 and it currently contains around 6000 records for primates alone. This definitely makes it the most comprehensive dataset of primate parasites in existence.
The GMPD sounds amazing…so what’s the problem?
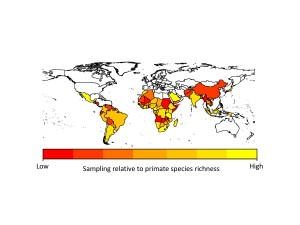
The problem with the GMPD (and this is a feature of virtually all datasets) is that there is sampling bias. Certain primates are sampled for parasites much more frequently than others. Chimpanzees, for example, are sampled for parasites all the time, whereas species such as tarsiers are sampled much less often. This has the effect of making it look like chimpanzees have far more parasites than tarsiers, simply because they have been sampled more often. In analyses using the database we usually deal with this problem by adding sampling effort into our models, so we give less emphasis to high numbers of parasites in primates we have lots of samples for. Unfortunately this problem is also evident when we look at parasites (things like malarial parasites are often sampled because of their importance to human health) and geographic regions (areas with primate research stations are sampled far more regularly than more remote regions). If we hope to use the GMPD data to make reliable predictions about future risks to humans, we need to identify gaps in our knowledge of primate parasites.
So what did you do?
Without going into the technical details, we looked across the primate phylogeny and primate geographic ranges to identify gaps in our knowledge, and used statistical models to investigate what factors led to primates and geographic areas being relatively well- or relatively poorly-sampled. We also used species accumulation curves to extrapolate parasite species richness for primates.
Where are the gaps in our knowledge?
We found that apes (chimpanzees, gorillas and orangutans) were generally better-sampled than other primates, but there was incredible variation in sampling among all other major primate groups. Apart from apes, the primates that researchers appear to sample most are the species they encounter most often, i.e., widespread, terrestrial, diurnal species. However, some primates were sampled more often because they are already intensively studied for other research, because they live in frequently visited field sites, or because of their importance in medical research. Across countries, we found that in general, parasite sampling is highest in countries with more primates to sample. We expected that the GDP of the countries would also affect sampling effort, with wealthier countries having more money for disease monitoring. However, we found no evidence for this in our analyses, probably because most research on primate diseases is not funded by the country in which the research takes place.
When we extrapolated parasite species richness values we found that even within our best-sampled primates and countries, we are missing a lot of parasites. On average we predicted that 38-79% more parasite species than currently reported in the GMPD should be found in our best sampled primate species, and 29-40% more parasite species than currently reported in the GMPD should be found in our best sampled countries. This emphasizes exactly how poor our sampling is across all primates and countries. Another concern is that although viruses make up only 12% of the parasites in our dataset, viruses arguably present the greatest zoonotic disease threat to humans because their fast rates of evolution should allow them to easily adapt to new hosts.
What next?
Identifying parasite sampling gaps across primate species and geographic regions is only the first step; we need to find strategies to minimize these sampling gaps if we are to predict which primate diseases may emerge in humans. One solution is to set research priorities based on the sampling gaps, for example, by focusing effort and funding on relatively poorly-sampled primate species, arboreal primates, those with small geographic ranges, or those found in relatively poorly-sampled regions of South East Asia, Central and Western Africa, and South America.
Focusing on relatively poorly-sampled primate species and areas may improve our general understanding of primate parasites, but it is only one factor in predicting risk to humans. For example, hosts are more likely to share parasites with their close relatives than with more distant relatives. Thus, continuing to focus our sampling efforts on parasites of our closest relatives (chimpanzees, gorillas and orangutans) may provide the greatest return in the case of risks to humans. This is particularly important because we found that chimpanzees are expected to have 33-50% more parasites than currently found in the GMPD. In addition, ecological similarities also influence parasite sharing among primates, and humans share more parasites with terrestrial than arboreal primate species. As with sampling effort, this probably reflects higher contact rates among humans and terrestrial primates compared to arboreal primates. As a related issue, a host living at higher density is expected to have higher prevalence of parasites and may have more contact with human populations or our domesticated animals, thus increasing opportunities for host shifts to humans. The large numbers of zoonotic emerging infectious diseases with rodent or domesticated animal sources also highlight the importance of rates of contact and host density for disease emergence in humans.
In conclusion Sampling effort for primate parasites is uneven and low. The sobering message is that we know little about even the best studied primates, and even less regarding the spatial and temporal distribution of parasitism within species. Much more sampling is needed if we hope to predict or prevent future emerging infectious diseases outbreaks.
Author
Natalie Cooper
nhcooper123
ncooper[at]tcd.ie
Photo credit
Natalie Cooper, wikimedia commons


